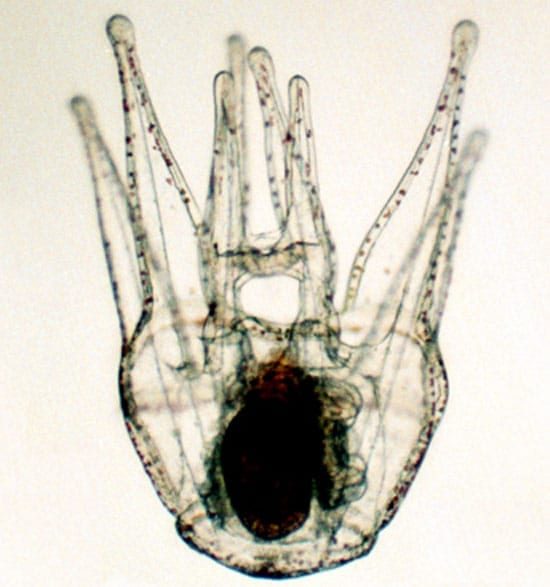
Sea Urchin Genome Yields New Understanding of “Chemical Defensome”
November 13, 2006
The Sea Urchin Genome Sequencing Consortium, a group of 240 researchers from more than 70 institutions in 11 countries, recently announced the sequencing of the California purple sea urchin, Strongylocentrotus purpuratus. Three biologists from the Woods Hole Oceanographic Institution (WHOI) were among those participating in describing the sea urchin genes. They helped to identify a large group of genes encoding proteins involved in protecting the sea urchin from toxic chemicals.
Reporting in the November 10 issue of the journal Science, the consortium researchers announced the high-quality “draft” sequence, covering more than 90 percent of the sea-urchin genome, and provided detailed descriptions of genes involved in a variety of biological processes ranging from immune function to sensory biology. Those descriptions appear in 41 papers in the December 1 issue of the journal Developmental Biology.
The sea urchin genome contains over 814 million letters, spelling out 23,300 genes. The WHOI scientists, in collaboration with researchers from Stanford University’s Hopkins Marine Station and several other institutions, identified over 400 of these genes that are part of the “chemical defensome,” an integrated network of genes and pathways that allow animals to mount an orchestrated defense against toxic chemicals.
The defensome genes include those for enzymes involved in biotransformation — the conversion of chemicals to less-toxic derivatives — as well as those encoding proteins involved in the expulsion of chemicals or their metabolites from cells. Also included in the defensome are genes for proteins called transcription factors, which sense the presence of toxic chemicals and turn on the genes for biotransformation enzymes and transporters.
The defensome characterization was led by Jed Goldstone, a postdoctoral scholar in the WHOI Biology Department, and Amro Hamdoun of Stanford University, and also included two WHOI senior scientists, biologists Mark Hahn and John Stegeman. For many years, the WHOI group has studied defensome genes in a variety of animals. The sea urchin research provides the first comprehensive, genome-wide assessment of the defensome in any animal. The sea urchin defensome is notable because several of the gene families in it have undergone expansion in the sea urchin lineage, suggesting an especially important role for chemical defense in these animals.
The overall sequencing project was led by Drs. Erica Sodergren, George Weinstock, and Richard Gibbs at the Human Genome Sequencing Center at Baylor College of Medicine, and Drs. Eric Davidson and Andrew Cameron of the California Institute of Technology.
Sea urchins are echinoderms (Greek for “spiny skin”), marine animals that originated over 540 million years ago and include starfish, brittle stars, sea lilies, and sea cucumbers. There was great interest in the sea urchin as a target for genome sequencing because these animals share a common ancestor with humans. That ancestor lived over 540 million years ago and gave rise to the Deuterostomes, the superphylum of animals that includes phyla such as echinoderms and chordates, the phylum to which humans and other vertebrates belong. All Deuterostomes are more closely related to each other than they are to any other animals not included in the Deuterostome superphylum. For example, among sequenced genomes, the sea urchin genome is closer to the human genome than to genomes of flies and worms.
“Each genome that we sequence brings new surprises. This analysis shows that sea urchins share substantially more genes and biological pathways with humans than previously suspected,” said Dr. Francis S. Collins, director of the National Human Genome Research Institute at the National Institutes of Health. “Comparing the genome of the sea urchin with that of the human and other model organisms will provide scientists with novel insights into the structure and function of our own genome, deepening our understanding of the human body in health and disease.”
The comparison of the genes of the sea urchin to the human gene list shows which human genes are likely to be recent innovations in human evolution and which are more ancient. It also shows which human genes have changed slowly in the lineage from the ancestral Deuterostome animal and which genes are evolving rapidly in response to natural selection.
Although, as invertebrates, sea urchins have a radically different morphology from humans and other vertebrates, their embryonic development displays basic similarities, an important shared property of Deuterostome animals. The evolutionary relationship to humans makes the sea urchin, with its many useful properties such as ease of isolation of eggs and sperm and transparent embryos, a valuable model to study the process of development and help understand human development. The development of the animal occurs through a complex network of genes, and the sea urchin is one of the main models for systems biology, the description of how the building blocks of an animal interact in time and space.
The chemical defensome described by the WHOI team and their colleagues may be especially important for protecting sea urchin embryos from toxic chemicals during embryonic development. Indeed, the team found that many of the defensome genes are expressed, or “turned on,” in embryos. In addition, based on comparisons with the previously sequenced genomes of other invertebrates and vertebrates, the researchers suggested that some defensome genes may have originally functioned as part of the complex network of developmental regulatory genes, and then later evolved their defensive roles.
The WHOI researchers were funded by grants from the National Institute of Environmental Health Sciences, at part of the National Institutes of Health.