Bacteria Exhibit Altruistic Behavior
Some microbes make antibiotics to benefit their kin
When it comes to bacteria protecting themselves, it’s all in the family.
A new study shows that marine bacteria can produce antibiotic compounds that kill unrelated bacteria but do not harm their closest relatives. What’s more, only some of the bacteria within their family make the antibiotics, so that the action of a few protects the entire group, according to scientists at Woods Hole Oceanographic Institution (WHOI) and the Massachusetts Institute of Technology (MIT).
“I think this is showing that bacteria can form social networks and have a partitioning of tasks,” said Tracy Mincer, a microbiologist at WHOI and co-author of the study, which was published in the Sept. 7 issue of Science. “Bacteria are coordinating with one another, and they’re sharing resources.”
The research team started working on the experiments about six years ago, when Mincer was a postdoctoral scientist at MIT and began collaborating with microbial ecologist Martin Polz. Their goals were twofold: to find previously unknown compounds made by bacteria that might have medical applications and to learn what functions those compounds accomplished for the bacteria in their natural environment.
“Even then, Martin was thinking about these networks of organisms and how they were teaming up with one another in some way,” Mincer recalled.
Keeping it in the family
Polz’s lab had amassed a collection of more than 20,000 strains of marine bacteria of the genus Vibrio. Most of the strains came from coastal areas of New England. Most are free-living microbes that feed on bits of organic matter in seawater. Many are associated with marine animals or plants. A few are disease-causing agents in fish or other sea animals.
Mincer helped design a procedure to screen each strain for its ability to kill other microbes. The test involves placing two strains of Vibrio on nutrient gel in the same petri dish and watching to see whether they both grow and thrive or one kills the other.
MIT student Sarah Proehl screened 185 strains, but the results were complex, with thousands of interactions between strains. They weren’t analyzed until postdoctoral scientist Otto Cordero arrived in Polz’s lab a few years later and began to use his programming skills to develop software to tease out what the bacteria were doing.
Cordero put the Vibrio strains into kin groups, or clades, based on the DNA sequences of six genes all the cells had. Then he mapped the antibiotic production, resistance, and vulnerability of each strain. Almost half (44 percent) of the strains produced an antibiotic compound that killed at least one other strain, with each strain producing a different antibiotic. Most of the strains (86 percent) were killed by at least one other.
The most notable finding was that each strain’s antibiotic killed only “outsiders.” Members of the strain’s own clade might be vulnerable to the antibiotic made by non-relatives, but they were always resistant to antibiotics made by their closest kin.
A valuable collection
The study demonstrated the value of Polz’s huge collection of Vibrio strains as potential sources of drugs and other useful compounds, Mincer said.
“Some of these vibrios are the fastest-growing microbes on the planet; they can settle and respond and grow very, very quickly,” said Mincer. “So it makes sense that they would also be able to defend their turf and attract their friends to make a more competitive biofilm and have a very stable community.”
A biofilm is a coherent, often slimy aggregation of related bacteria. Biofilms can thrive where individual bacteria would struggle or not survive at all. That helps explain why a bacterium would invest a lot of energy in producing a compound that benefits other bacteria as well as itself, and why a specific environment selects for such traits.
“In a lot of ways, this is altruistic behavior,” Mincer said. “They make this molecule at a significant metabolic expense, for the betterment of the group.”
Super-killers and jumping genes
The researchers dubbed a few strains of Vibrio“super-killers” because their antibiotics killed the cells of at least ten other strains. To find out more about the antibiotic made by one super-killer, Cordero and fellow MIT postdoctoral scientist Hans Wildschutte subjected the super-killer cells to a process that interrupted various genes and then looked for “knockouts,” cells that were no longer able to make the antibiotic. In this way, they discovered which genes had been involved in making the antibiotic.
They found that the super-killer’s antibiotic was a previously unknown compound made by a biochemical pathway that had never been observed before. They also found that the genes to make the antibiotic are “jumping genes,” that is, they can be clipped out of the chromosome and inserted into different locations in the bacteria’s DNA, either in the same or in different bacteria.
Where the genes came from and why only certain organisms acquired them is not well understood. The genes could have transferred from other bacteria or viruses. They might even have been floating free in seawater, available for pickup by any bacteria able to do it.
Of course, an acquisitive bacterium that brings in the genes to make an antibiotic runs a risk: If it starts making an antibiotic that it is not naturally resistant to, it will kill itself. But if the acquired genes are beneficial, “you’ve got an organism that has taken upon itself to start to collect these genes, and that becomes advantageous,” Mincer said. “The organism starts to form a community around it—an entourage, if you will, of other cells.” Then the ability to make the antibiotic becomes useful to the entire community.
Next steps
Mincer thinks strains of bacteria that acquired one set of genes may be more likely to pick up others as well, and that has big implications for the search for useful compounds: With thousands of cell stocks to survey, a scientist’s time might be better spent delving deeper into the cupboard of compounds produced by a strain that has already yielded one valuable product than looking for candidate molecules from completely different strains.
Mincer said going back over the same ground to find new drugs reminded him of how gold mining was done in the early 1900’s where he grew up in Alaska. Prospectors there used floating dredges that chewed up the riverbanks and sifted the rocky material to look for gold.
“The dredges were geared to get gold nuggets that were very small, up to about fingertip size, and they just threw the tailings out the back,” he recalled. “Now people have gone back to where these gravel areas are and turned over the rocks and found gold nuggets the size of potatoes. It turns out they just didn’t have the sieving sizes right for the really big stuff.”
In addition to looking for new compounds of interest, Mincer is also studying some of the compounds the group already found. He has partially analyzed one super-killer’s antibiotic and found that it is a hybrid molecule, part peptide and part fatty acid. He thinks it may be a porin, a type of molecule that makes the outer membrane of susceptible cells leaky, essentially making the cells fall apart.
He and his colleagues are also trying to understand the role played by this and other antibiotics in Vibrio’s natural environment. In addition to killing competitors, an antibiotic may signal to closely related cells that this is a safe place to settle and form a stable community. The researchers also hope to learn more about what, if anything, other members of the community—those that benefit from having an antibiotic-producing cousin—contribute to the group.
This research was supported by the National Science Foundation, the Gordon and Betty Moore Foundation, the Broad Institute, the Netherlands Organisation for Scientific Research, and the Woods Hole Center for Oceans and Human Health.
Slideshow

Slideshow
 Microbiologist Tracy Mincer of WHOI is working to discover compounds made by marine microbes that could have applications in human medicine. In a study recently published in the journal Science, he and his colleagues identified and partially characterized several potent antibiotics. (Photo by Tom Kleindinst, Woods Hole Oceanographic Institution)
Microbiologist Tracy Mincer of WHOI is working to discover compounds made by marine microbes that could have applications in human medicine. In a study recently published in the journal Science, he and his colleagues identified and partially characterized several potent antibiotics. (Photo by Tom Kleindinst, Woods Hole Oceanographic Institution)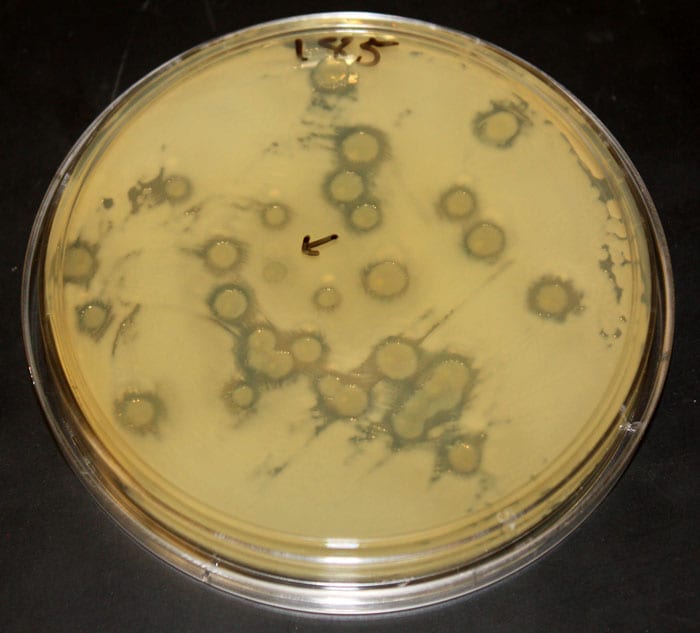 A test plate shows the killing effect of some strains of Vibrio bacteria on another strain. The nutrient gel in this dish was evenly covered with a yellowish "target" strain. Then small spots of other bacterial strains were placed at random on top. "Halos" indicate where added strains killed the target bacteria. The size of a halo is related to the number of cells that died. The arrow indicates a spot of added bacteria that lacks a halo; the strain in this spot did not kill the target strain. (Photo courtesy of Hans Wildschutte)
A test plate shows the killing effect of some strains of Vibrio bacteria on another strain. The nutrient gel in this dish was evenly covered with a yellowish "target" strain. Then small spots of other bacterial strains were placed at random on top. "Halos" indicate where added strains killed the target bacteria. The size of a halo is related to the number of cells that died. The arrow indicates a spot of added bacteria that lacks a halo; the strain in this spot did not kill the target strain. (Photo courtesy of Hans Wildschutte)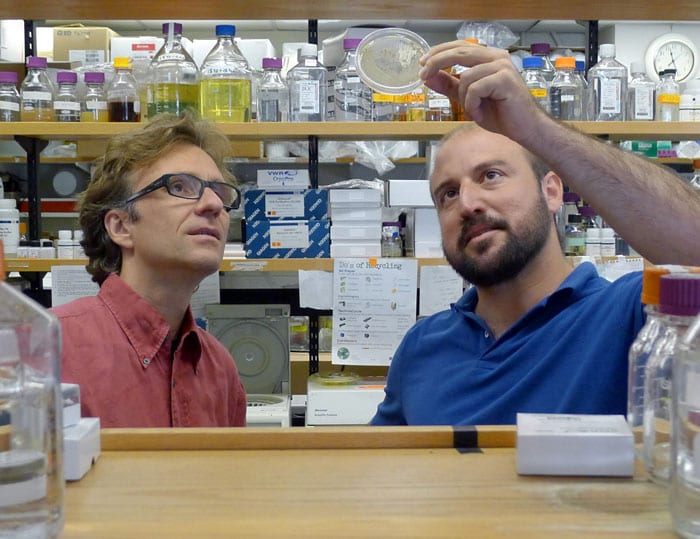 Microbial ecologist Martin Polz (left) and theoretical biologist Otto Cordero of MIT led a team investigating the interactions among 185 strains of the marine microbe Vibrio. They found that many of the strains produce antibiotics that don't kill strains that are closely related to themselves, but do kill other strains that are not closely related to them. (Photo by James Long, Massachusetts Institute of Technology)
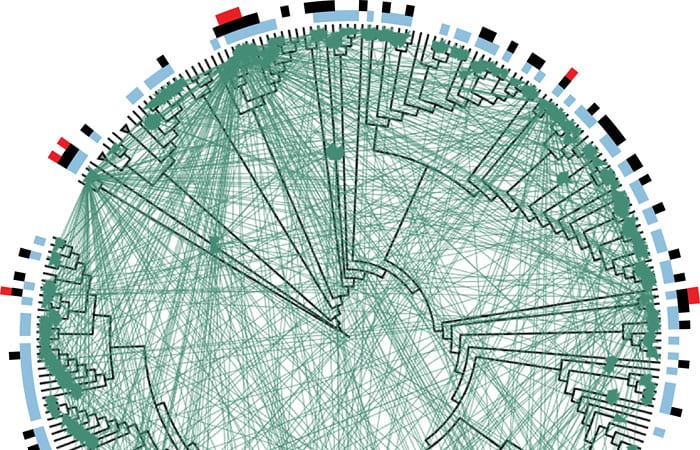
Microbial ecologist Martin Polz (left) and theoretical biologist Otto Cordero of MIT led a team investigating the interactions among 185 strains of the marine microbe Vibrio. They found that many of the strains produce antibiotics that don't kill strains that are closely related to themselves, but do kill other strains that are not closely related to them. (Photo by James Long, Massachusetts Institute of Technology)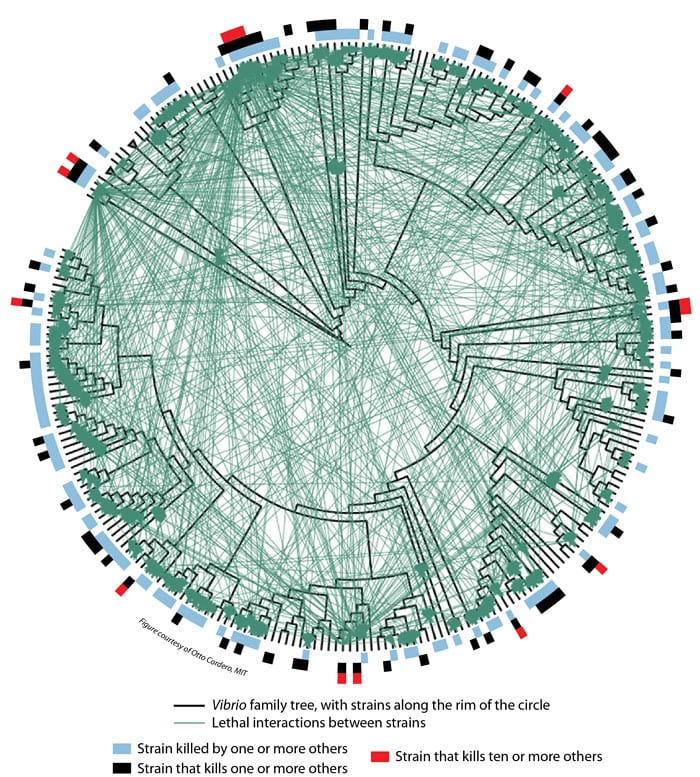 Researchers created this diagram to summarize their findings about lethal interactions among 185 strains of Vibrio bacteria. Individual strains of bacteria are represented by the black lines protruding from the rim of the circle. Those adjacent to each other on the rim are more closely related than those farther apart. The black lines radiate from a Vibrio family tree (displayed in circular form rather than the more common vertical "tree" arrangement). Each green line represents an interaction between two strains in which one of the strains produces an antibiotic that kills the other. Blue bars along the circumference indicate strains that are killed by at least one other strain. Black bars indicate strains that kill at least one other strain. Red bars indicate "super killers"—strains that kill at least ten other strains. (Otto Cordero, Massachusetts Institute of Technology)
Researchers created this diagram to summarize their findings about lethal interactions among 185 strains of Vibrio bacteria. Individual strains of bacteria are represented by the black lines protruding from the rim of the circle. Those adjacent to each other on the rim are more closely related than those farther apart. The black lines radiate from a Vibrio family tree (displayed in circular form rather than the more common vertical "tree" arrangement). Each green line represents an interaction between two strains in which one of the strains produces an antibiotic that kills the other. Blue bars along the circumference indicate strains that are killed by at least one other strain. Black bars indicate strains that kill at least one other strain. Red bars indicate "super killers"—strains that kill at least ten other strains. (Otto Cordero, Massachusetts Institute of Technology)
Related Articles
- 4 Potential Solutions for Corals in Crisis
- An ocean of opportunity
- Putting the ‘nuclear coffin’ in perspective
- Chasing Ocean ‘Snowflakes’
- Journey to the Bottom of the Sea
- Life at the Edge
- The Discovery of Hydrothermal Vents
- Mission to the Ocean Twilight Zone
- What Happened to Deepwater Horizon Oil?
- New Device Reveals What Ocean Microbes Do
- Minerals Made by Microbes
- Two Chemical Roads Diverge in an Open Ocean
- A Mighty Mysterious Molecule
- Earth’s Riverine Bloodstream
- Short-circuiting the Biological Pump
Featured Researchers
See Also
- Natural Antibiotics from Microbes News release from MIT media office
