Recipes for Antibiotic Resistance
A student examines the bacterial bouillabaisse in the coastal ocean
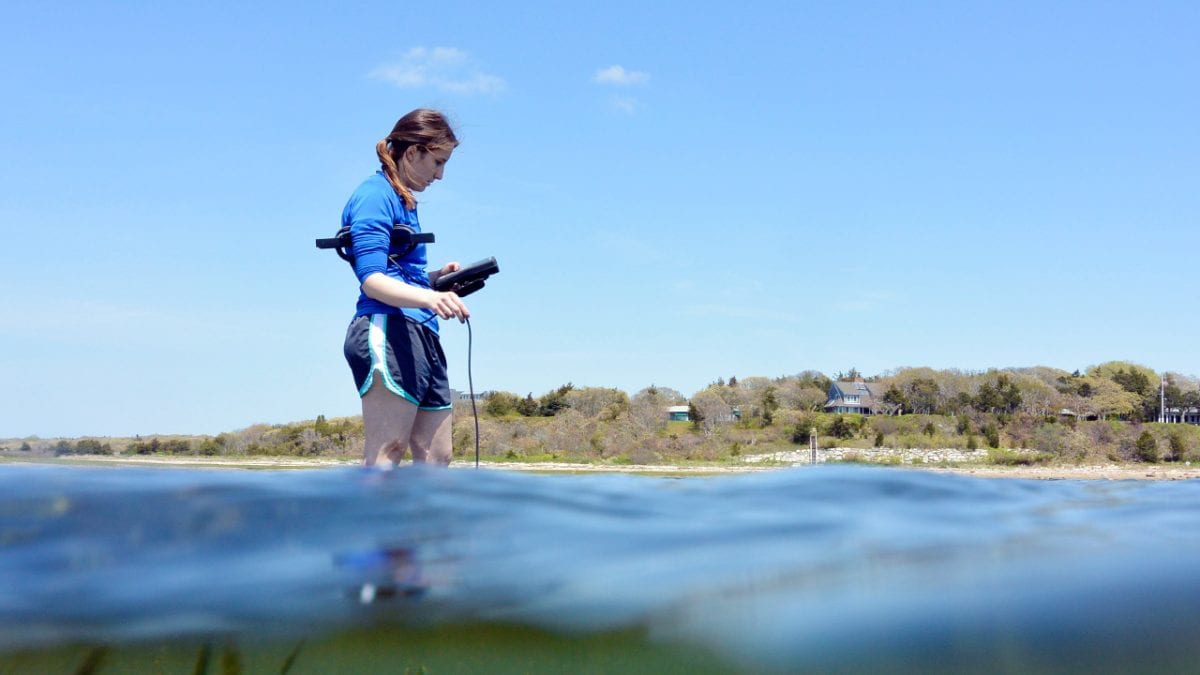
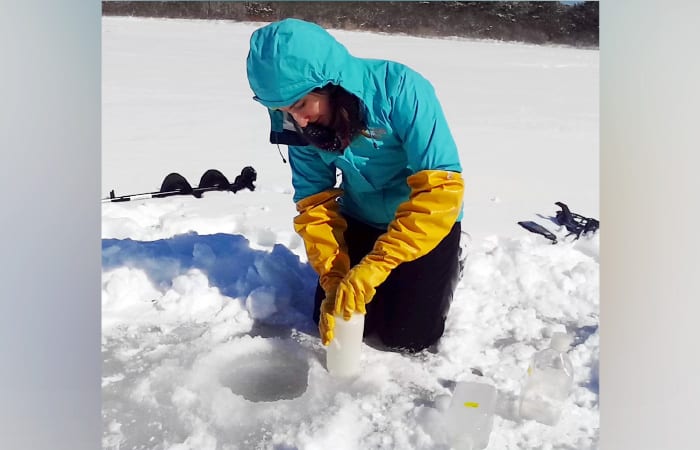
As they trekked through a thick layer of snow, Megan May and her Ph.D. advisor Rebecca Gast lugged their water sampling equipment down an unplowed access road toward a frozen coastline. They drilled through the ice, fruitlessly hitting sand in their first few attempts, until they finally struck water. All this effort was not to collect the water itself, but the invisible life suspended within it.
This invisible life exists in a vast microscopic realm, where microbes compete for limited resources. They have developed chemicals to perform a variety of functions, such as signaling other cells, stimulating metabolic functions, and occasionally warding off competitors.
Over the past century, humans have learned to use some of these bacterial chemicals to create antibiotics that fight infections. However, the Centers for Disease Control and Prevention (CDC) reported a rising trend in recent years of some bacteria developing resistance to antibiotics. When bacteria develop resistance faster than scientists can develop new antibiotics, infections that were once easy to treat become life-threatening.
A 2013 report by the CDC stated that antibiotic resistance in bacteria is one of our most serious health threats. More than two million people per year become sick with antibiotic-resistant infections, with at least 23,000 dying as a result. These infections require longer hospital stays, costlier and/or longer treatments, and additional doctor visits and health care. They cause greater disability compared to infections easily treated with antibiotics and up to $20 billion in excess direct health-care costs, the CDC reported.
To combat this growing problem of resistance, scientists need to know what they’re fighting against. This is where May’s research comes in.
The oceans contain an untold number of species of bacteria, constantly mixing, feeding, reproducing, and exchanging genes in a chaotic milieu. This setting allows May to investigate when and how bacteria naturally produce antibiotic chemicals and develop resistance to them.
In addition, the oceans are rapidly changing. The skyrocketing use of antibiotics in health care and agriculture has led to an influx of antibiotic residues, resistant bacteria, and resistance-coding genes into the oceans. Is human activity significantly disturbing natural processes? Could an accelerating proliferation of resistant bacteria pose a threat to people?
Eye on an invisible world
May first became interested in microbes as part of a seventh-grade science experiment. To explore the effects of hand washing, she cultured bacteria found on hands.
“I ended up having these petri dishes in our house and growing bacteria above the fridge because that was a nice warm place,” May said. “In retrospect, I wouldn’t do that now—not that safe—but it was awesome that my parents let me do that.”
“I really got excited about seeing these little things that are all around us, but are invisible otherwise,” she said. “I was kind of a nerd and went to various math and science summer courses that also developed my interest.”
As an undergraduate at DePauw University, May examined the microbial diversity of the DePauw Nature Park ponds. She also interned at Mount Desert Island Biological Laboratory in Maine, worked at the University of Tennessee to study bacteria in deep marine sediments, and spent a semester in Ecuador studying ecological sanitation problems. All these experiences helped her develop what she describes as a “Venn diagram” of intersecting interests: aquatic environments, microbiology, and public health.
May learned about Gast’s research, which had found evidence of resistant bacteria in marine animals. It fit all the criteria of her Venn diagram, and she seized an opportunity to work with Gast as a graduate student in the MIT-WHOI Joint Program. They expanded the research from looking at resistance in microbial life in animals to resistance in microbial life in the water and sands.
Antibiotics on the rise
Bacteria have been using chemicals to communicate with one another for millennia. Consequently, some bacteria have evolved methods of resistance to these chemicals’ effects, allowing them to ignore or modify chemical signals.
Scientists learned to repurpose these chemicals as weapons against bacterial infections, creating antibiotics. For example, they found that normally helpful compounds made by bacteria in the soil or ocean can be deadly to pathogenic bacteria in animals that had never encountered these compounds before. While antibiotics undoubtedly have saved countless lives and greatly advanced food production, their overuse is now creating a serious threat.
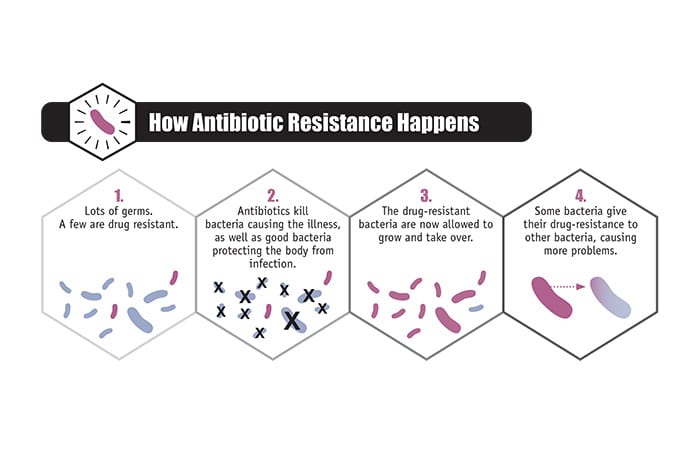
The human body naturally contains trillions of bacteria. Some are beneficial and necessary to maintain a healthy body, some are neutral, and some are harmful. Antibiotics that combat harmful bacteria flood our bodies and kill off good and infectious bacteria alike. This creates a “selecting environment,” as May put it. Normally, the body contains some bacteria with resistance to a given antibiotic. The antibiotic kills off non-resistant bacteria, allowing resistant bacteria to thrive and ultimately increasing their number.
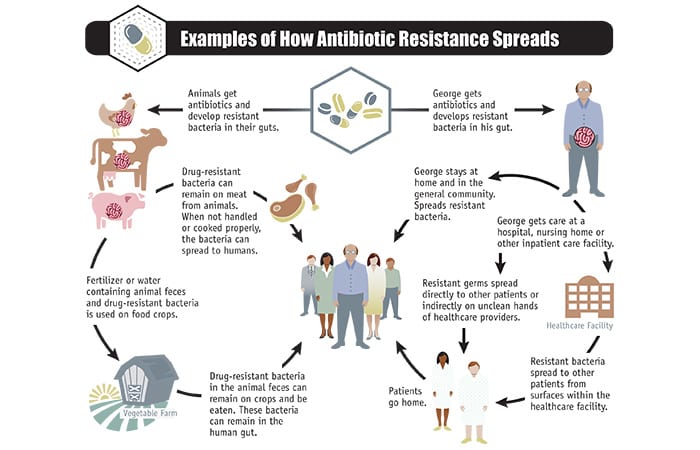
“We use antibiotics a lot,” says May, “and not just for human health.” Antibiotics are also commonly used to promote growth and prevent infections in livestock in agriculture and aquaculture. Long-term exposure to antibiotics allows resistant bacteria to accumulate within animals. These bacteria can remain on meat harvested from these animals and can spread to humans. Resistant bacteria in animals can spread in fertilizer or water containing the animals’ feces. They can contaminate crops and leach into soils, allowing resistant bacteria to diffuse throughout the ecosystem.
Changes in the coastal ocean
Similar principles hold true when antibiotic residues enter the environment. Normally, the environment contains many thousands of bacteria, some of which may have resistance to a given antibiotic. Residues from antibiotics can kill off non-resistant bacteria, giving resistant bacteria a competitive edge and increasing their number in the environment.
This process is complicated by the fact that bacteria can exchange genes with each other. “Bacteria are promiscuous,” as May put it. They can make connections and exchange genetic material often and quickly. Unlike humans, bacteria can incorporate others’ genes and express new traits immediately. It would be as if a blond person had sex with a red-haired person and gained the ability to grow red hair.
“A combination of many things are likely helping to increase resistance in the environment—antibiotic residues, antibiotic-resistant bacteria, naturally occurring resistance genes, and bacteria from human activities that normally wouldn’t be found in the environment,” May said.
“These all can intermix in the environment,” she said. “For example, antibiotic-resistant bacteria that normally live in the marine environment could transfer their resistance to a non-resistant pathogenic bacterium from human waste. That would create an antibiotic-resistant pathogen in nature, and that pathogen could potentially be transferred to humans who come in contact with it in the environment or consume food contaminated with it.”
Understanding resistance
May’s research centers on two questions:
- How, where, and when does resistance occur in the marine environment?
- Do pollution and other human activities affect patterns of resistance among marine microbes?
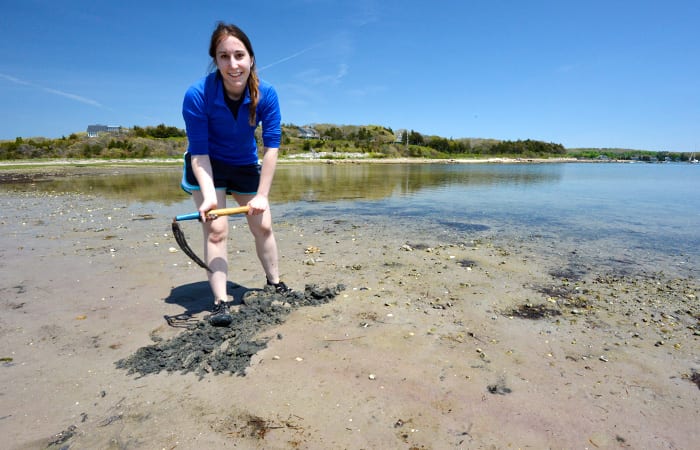
To answer these questions, May has been studying bacteria from six different field sites around Cape Cod. Over a year, across different seasons, she gathered samples of water and wet and dry sand from each site.
May grows three different kinds of bacteria found in her sand and water samples: Enterococcus, Vibrio, and “general marine bacteria.” Enterococcus often comes from feces and can cause disease in humans. Vibrio bacteria, some of which are pathogenic, are widespread throughout the marine environment. “General marine bacteria” are any kind of bacteria that can grow in a medium that resembles the composition of seawater.
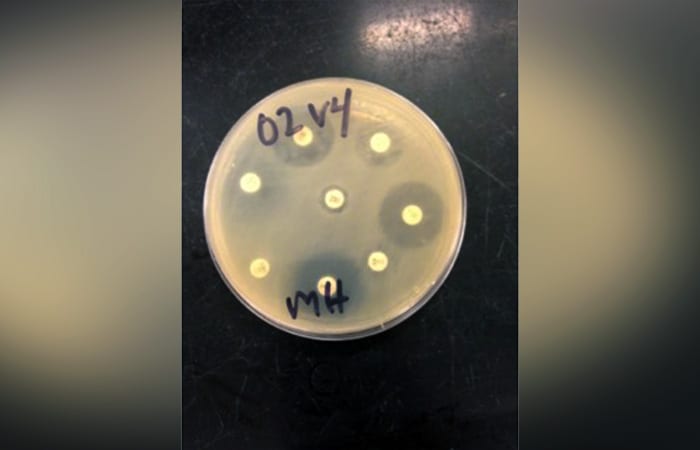
Into her bacterial cultures, she then inserts disks with known concentrations of antibiotic. She observes whether the chemical kills the bacteria, slows their growth, or doesn’t affect them at all—indicating whether particular bacteria are expressing resistance to a given antibiotic.
May selected six Cape Cod study sites, with comparatively high, mild, or low levels of human activity and pollution. Comparing results from different areas will give May clues about whether human activity is affecting patterns of resistance.
The phase of May’s research that required her to trudge to frozen coastlines has transitioned into the even more painstaking work of culturing, analyzing, and organizing copious samples.
“I figure, ultimately, you’re spending all this time on a given project, you want to be excited about it, and sometimes it can be hard to be excited about something, when you’re working late at night or when your experiment went wrong another time, or whatever, so there’s got to be some kind of internal passion to it,” she said. “I want my information to be relevant and useful.”
The results of her research, she hopes, will be meaningiful for microbiologists, oceanographers, health-care policymakers, coastal resource managers, tourists, and even seventh-graders looking for an inspiring science fair project.
This research was funded by a National Science Foundation Graduate Research Fellowship, WHOI’s Coastal Ocean Institute, WHOI’s Ocean Ventures Fund, WHOI’s Grassle Family Foundation, and WHOI’s Academic Programs Office.
Resisting Resistance
Over the millennia bacteria have developed an array of chemicals that they require to function—to make their cellular machinery work, signal other cells, and occasionally ward off competitors. People have harnessed some of these chemicals to make antibiotics that cure infections.
In recent years, however, infectious diseases are rapidly becoming resistant to formerly effective antibiotics. Scientists are diligently working to give humanity the upper hand with new antibiotics and better diagnostic tests, but they cannot win the race alone.
Much of the problem stems form a vicious cycle: As people use more antibiotics, more antibiotic residues, resistant bacteria, and resistance-coding genes end up in the environment; As more bacteria are exposed to antibiotic residues and resistant bacteria or genes, more bacteria develop resistance.
Megan May, a graduate student in the MIT-WHOI Joint Program, is studying antibiotic resistance, and she suggested several ways that individuals can help in this fight.
“One thing I think is important is for people to understand is the difference between using antibiotics for bacterial infections and using them for viral infections,” she said.
Antibiotics work by targeting specific structures in bacteria that are not present in viruses, making antibiotics entirely ineffective against viral infections. According to the U.S. Centers for Disease Control, “up to 50 percent of all the antibiotics prescribed are not needed or are not optimally effective as prescribed.” May said that reducing the unnecessary use of antibiotics will reduce bacteria’s exposure to antibiotics and slow the development of antibiotic resistance.
May also recommends that when people do take antibiotics, they take the full course. “A lot of the time, your antibiotics are supposed to go for seven days, and you feel better after day four, so you just stop taking them. But that also promotes growth of resistant bacteria and doesn’t allow your infection to fully get better.”
People have also greatly increased their use of antibiotics in agriculture and aquaculture. To combat this trend, May recommends purchasing organic products.
“If you have the choice and you’re able to speak up and make change and say you want organic produce or meat, that also lessens the amount of antibiotics getting put into those products,” she said.
Supporting businesses that eliminate unnecessary use of antibiotics in their products encourages those practices and increases market demand for products with fewer antibiotics.
The development of resistance is a natural evolutionary process, so it is impossible to stop it completely. However, it can be slowed significantly by reducing the amount of antibiotics used needlessly in health care and food production.
—By Ashley Junger
Slideshow

Slideshow
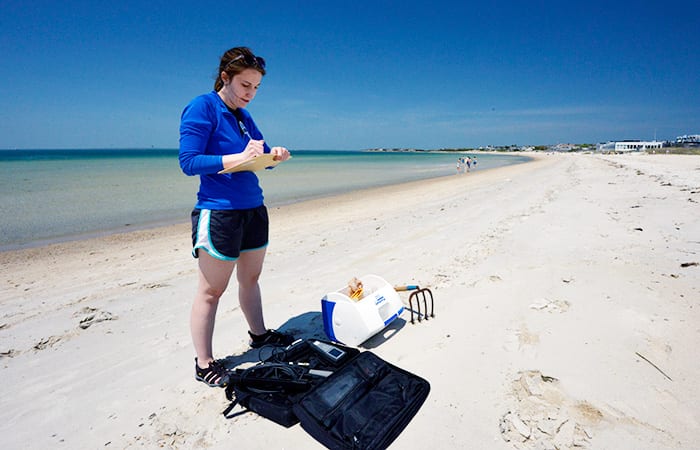
- For her research, Megan May, a Ph.D. student in the MIT-WHOI Joint Program, packs her equipment to collect samples of bacteria in seawater and dry and wet sand at Cape Cod beaches. (Ken Kostel, Woods Hole Oceaongraphic Institution)
- Graduate student Megan May samples throughout the year to see how bacterial populations change with seasons. She drills through frozen coastal waters to extract a water sample from one of her six study sites on Cape Cod. She will analyze bacteria in the samples to identify antibiotic-resistant genes present in the environment. (Rebecca Gast, Woods Hole Oceanographic Institution)
- Megan May cultures bacteria from her samples on petri dishes and then exposes them to antibiotic disks. Each white circle contains an antibiotic. The circles around the disks are known as zones of inhibition, the area where bacterial growth is inhibited by the presence of the antibiotic. The size of these zones provides information about how effective the antibiotic is at killing the bacteria, revealing the bacteria's degree of resistance. (Megan May, Woods Hole Oceanographic Institutioin)
- (The U.S. Centers for Disease Control)
- (The U.S. Centers for Disease Control)
Related Articles
- Cold, quiet, and carbon-rich: Investigating winter wetlands
- The 10,000-foot view
- A cascade of life
- An introduction to marsh bothering
- A toxic double whammy for sea anemones
- Squid Games
- Waves of inspiration
- Burrows on the beach
- Where the Rivers Meet the Sea
- Marshes, Mosquitoes, and Sea Level Rise
- The Unseen World on Coral Reefs
- The Recipe for a Harmful Algal Bloom
- Life at the Edge
- The Bacteria on Your Beaches
- Reassessing Guidelines for Oil Spill Cleanups
Featured Researchers
See Also
- Antibiotics Resistance Threats in the United States, 2013 Centers for Disease Control and Prevention 2013 report
- Rebecca Gast
- Sea Life Is Accumulating Pathogens Oceanus magazine
- Legions of Legionella Bacteria Oceanus magazine